BrightHub: Ingesting daily data files
Welcome to part 3 of our 5-part series on BrightHub, BrightWind’s wind resource data platform. In the previous article, we explained the data model used by BrightHub to store the manually entered sensor meta data. We will be borrowing a lot of terminology introduced in that article here, so if you haven’t already read it, check out BrightHub : Handling difficult wind resource data.
In this article, we will be exploring the file ingestion system we use to extract met mast metadata from the daily data files and how we combined this ingestion system with the data model we introduced in the previous article.
What do we want from our file ingestion system?
We outlined in the previous article a data model which could be used to handle all possible metadata changes that can happen to sensors mounted on met masts.
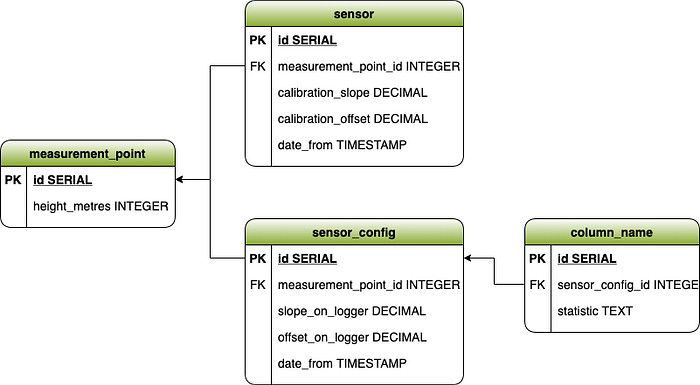
While the measurement_point and sensor tables are filled in manually by an analyst, manually filling in the sensor_config and column_name tables in particular can be rather tedious. We wanted a file ingestion system that would extract slope and offset values programmed into the logger from each data file as well as extracting the measurement data of each sensor.
The file ingestion system needed to capture how the values changed over time, seeing as we are interested in the entire measurement period of a sensor, not just the most recent values. It would also need to integrate seamlessly with our pre-existing data model, so that the metadata entered by the analyst could be linked to the extracted metadata. Finally, it needed to be able to produce the same results regardless of the order the files were being processed in.
Load and compile
We were lucky enough to work with senior consultants from fourTheorem to help us come up with a solution. They suggested implementing a load and compile system to handle our file ingestion. This entailed writing metadata to loading tables as each daily file is being ingested and then compiling the metadata in these loading tables to a set of compile tables. After the compile process was complete, the entries in the compile tables would be a representation of how our metadata changed over time. Furthermore, provided that we used primary keys that would always be reproducible each time we load in daily files, we could ensure idempotency of the system. This meant that we could safely link the manually entered metadata in our data model and the measurement data in our time series database to the metadata extracted by the file ingestion system.
In order to support this load and compile system, we needed to make some changes to our database schema:
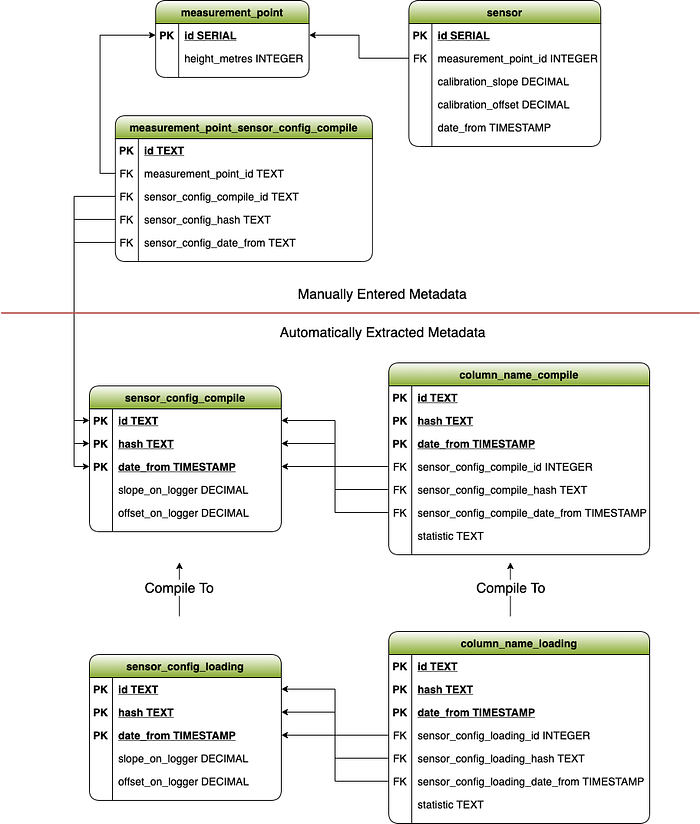
The main changes to the schema were as follows:
- We broke out the sensor_config and column_name tables into loading and compile tables
- On each of these tables, the integer id was replaced with a text id. We needed an id that would be the same each time we load in the same data, so we could safely tie our metadata to it. Auto-incremented integers wouldn’t give us this reproducibility.
- We added a hash field. This is a hashed value of the row itself, excluding the date_from field. We will use this in the compile process to quickly determine whether a config has changed or not
Now that our database was ready, we could start processing the files. As each file was being processed, our file-ingestion system would extract the metadata from the file and load it into the loading tables with the corresponding values.
For example, if the data file contained metadata for a sensor config with a slope_on_logger value of 0.1 and an offset_on_logger value of 0.2, then a row would be added to the sensor_config_loading table with the following values:
- id would be a string combination of the logger serial number and the name of the sensor config in the file. This combination would be a universally unique value and would be the same no matter how many times we load in that same data file
- hash would be a hash of the configuration values (slope and offset in this case). Seeing as our file processing logic is written in Python, we simply hashed a json representation of our slope and offset values
- date_from would be the first timestamp in the daily data file
- slope and offset would be the corresponding slope and offset values in the metadata section of the daily data file
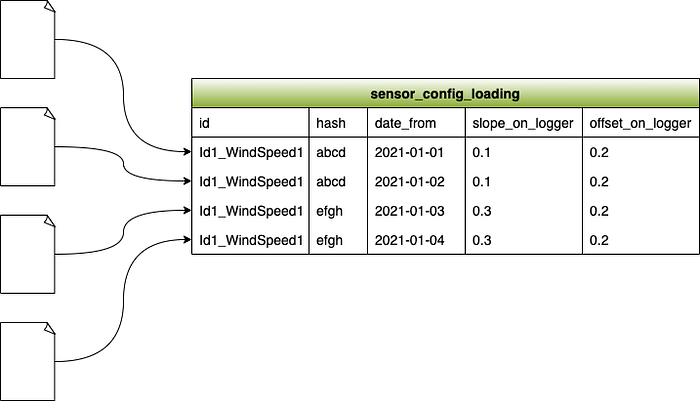
When we load the next file in, another row gets added to the sensor_config_loading table for that sensor. If the slope_on_logger and offset_on_logger hasn’t changed, then the row will be the exact same apart from the date_from.

Continuing with our example, lets imagine the third file we load in has a slope_on_logger value of 0.3, meaning that its configuration has changed. For this file, the row added into the sensor_config_loading table would have a different slope_on_logger value and therefore a different hash value than the previous two files. Any subsequent files that contain that same configuration change would generate the same hash value.
Next we want to compile the sensor_config_loading table into the sensor_config_compile table, which will represent how the configuration changed over time. To do this we use a SQL query to do the the following:
- Select all rows in our sensor_config_loading table that have the same id. This shows us all configurations for a given sensor over time
- Group these rows by their hash value. This puts all sensor configuration snapshots with the same configuration (i.e. the same slope_on_logger and offset_on_logger) in the same group
- Select the minimum date_from of all of the snapshots in each group. This is the first date where the new sensor configuration was observed
- Add an entry to the sensor_config_compile table that reflects this configuration and make sure to write the minimum date_from to represent the start date of this configuration
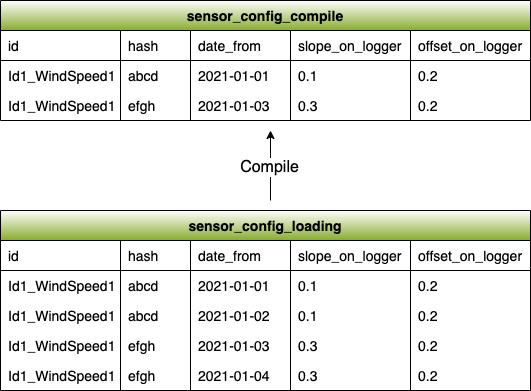
We repeat the same process for every sensor_config and column_name in every daily data file and we get a full representation of all configuration changes over time for an entire met mast. The analyst can then link their manually entered metadata to the extracted metadata.
Next Steps
Now that we had a data model which would handle all sensor metadata scenarios, and a file ingestion system that would extract all meaningful metadata from daily files, we could turn our attention to the sensor time series data. Our next article will discuss how we experimented with various time series databases to find one that could scale to handle our very spikey workflow, while also keeping to our relatively modest budget.
